Niklas Abraham
Machine Learning Student @ University of Tübingen

I am an MSc student in Machine Learning at the University of Tübingen (current grade average: 1.0), where my studies are embedded within one of Europe’s leading ML research hubs through links with the Max Planck Institutes and Cyber Valley. My coursework focuses on deep learning, statistical machine learning, and probabilistic inference. I completed my BSc in Simulation Technology at the University of Stuttgart in 2025 (grade 1.9, 5th in cohort of 22), with an elected focus on AI and bioinformatics. I am a scholarship holder of the German Scholarship Foundation (Studienstiftung des Deutschen Volkes).
My research interests center on advanced generative modelling (score-based diffusion, flow-matching), non-convex optimisation in deep learning, and reinforcement learning for sequential decision-making, with applications in structural biology and bioinformatics. Most recently, our work on converting diffusion-based co-folding models to deterministic probability-flow ODEs was accepted as a poster at the ICLR 2026 Workshop on Learning Meaningful Representations of Life, achieving a 20x sampling speed-up on FoldBench.
Previously, I worked as a research student at the Institute of Biochemistry and Technical Biochemistry (University of Stuttgart) under the supervision of Prof. Jürgen Pleiss, where I developed machine-learning frameworks integrating large protein language models (ESM-2, ESM-C) with curated biological databases to predict β-lactamase resistance phenotypes, and contributed to the Python Enzyme Engineering Database (PyEED). In October 2025, I won the Speed-Up Challenge at the Merck M-Boltz Hackathon by developing a fast deterministic flow-matching framework for accelerated protein structure generation. Starting April 2026, I will join Merck KGaA’s Group Science & Technology Office for a six-month internship.
Outside my academic work, I train for triathlon (next up: a 70.3 Ironman), coach climbing for children at KiSS e.V. twice a week, and read as much non-fiction as I can find.
News
| Apr 1, 2026 | Starting my six-month internship at Merck KGaA’s Group Science & Technology Office in Darmstadt. I will be working on metagenomic sequencing pipelines and benchmarking protein language models and DNA transformers for variant effect prediction. |
|---|---|
| Feb 1, 2026 | Our paper “Converting diffusions to flows accelerates sampling and suggests over-conditioning of co-folding models on sequence” has been accepted as a poster at the ICLR 2026 Workshop on Learning Meaningful Representations of Life (LMRL). |
| Oct 15, 2025 | Won the Speed-Up Challenge at the Merck M-Boltz Hackathon in Darmstadt by developing “Boltz Flow Matching”, a fast deterministic flow-matching framework enabling analytical conversion from score-based diffusion to flow equations for accelerated protein structure generation. |
| Oct 1, 2025 | Started my MSc in Machine Learning at the University of Tübingen. Looking forward to diving deeper into generative modelling, probabilistic inference, and reinforcement learning within the Cyber Valley research ecosystem. |
| Sep 15, 2025 | Completed my BSc in Simulation Technology at the University of Stuttgart (grade 1.9, 5th in cohort of 22). My thesis on predicting β-lactamase resistance phenotypes from sequence using protein language models is now listed on the Research page. |
Selected Publications
-
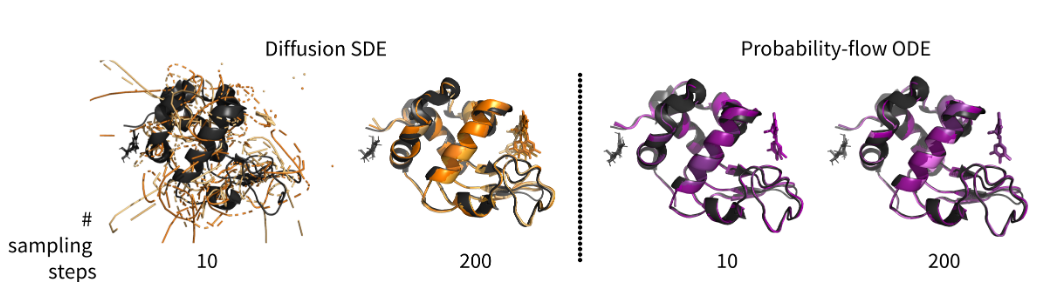 Converting diffusions to flows accelerates sampling and suggests over-conditioning of co-folding models on sequenceIn ICLR 2026 Workshop on Learning Meaningful Representations of Life (LMRL) 2026
Converting diffusions to flows accelerates sampling and suggests over-conditioning of co-folding models on sequenceIn ICLR 2026 Workshop on Learning Meaningful Representations of Life (LMRL) 2026 -
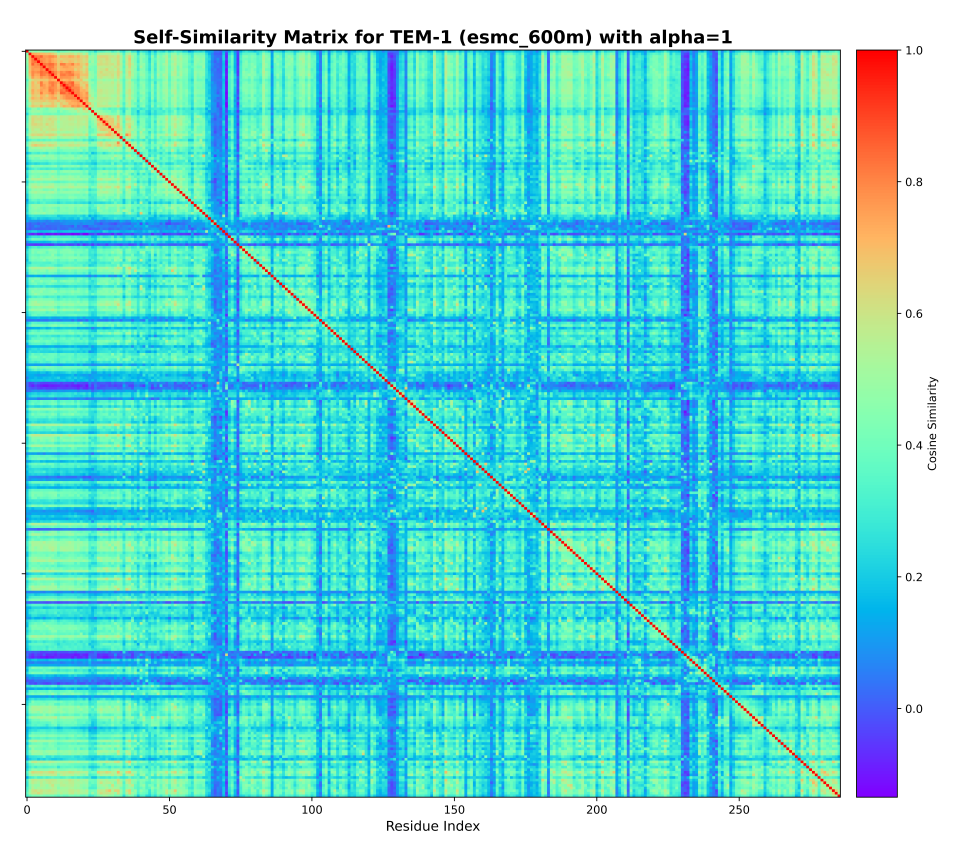
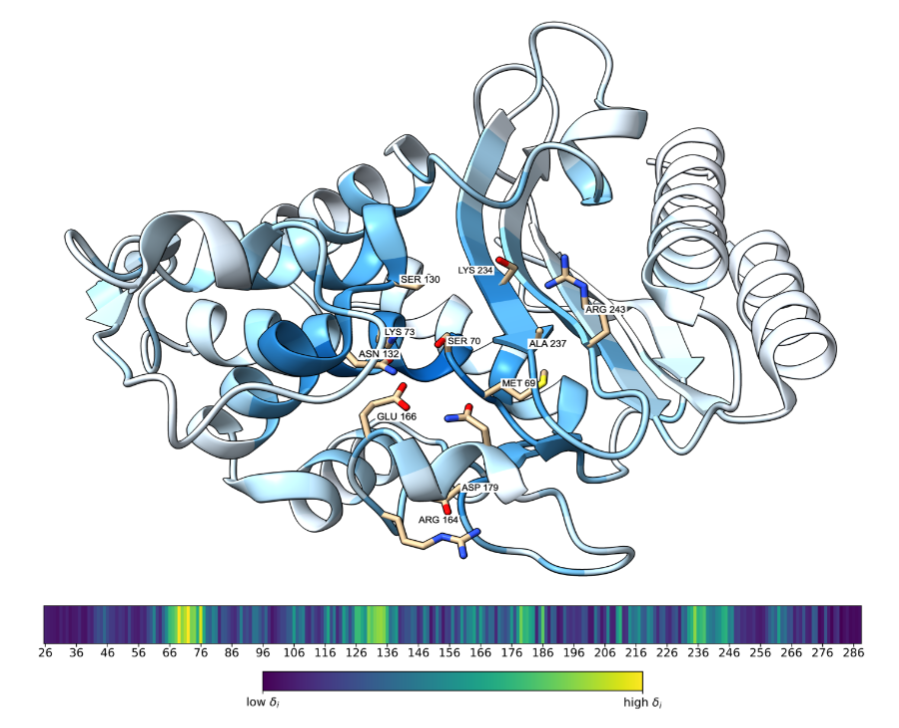 Geometric encoding of enzymatic mechanism in protein language model representationsManuscript in preparation 2026
Geometric encoding of enzymatic mechanism in protein language model representationsManuscript in preparation 2026